Infering Pathway or Transcript Factors activity from EnrichGT meta-gene modules
c_infreg.Rmd
library(EnrichGT)
#> Loading required package: dplyr
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
#> Loading required package: tibble
#> Loading required package: gt
#> Loading required package: cli
#>
#> ── EnrichGT ────────────────────────────────────────────────────────────────────
#> ℹ View your enrichment result by entring `EnrichGT(result)`
#> → by Zhiming Ye, https://github.com/ZhimingYe/EnrichGTDatabases
This is inspired from Saez-Rodriguez Lab’s decoupleR
package, https://saezlab.github.io/decoupleR/.
PROGENy is a comprehensive resource containing a curated collection of pathways and their target genes, with weights for each interaction.
CollecTRI is a comprehensive resource containing a curated collection of TFs and their transcriptional targets compiled from 12 different resources. This collection provides an increased coverage of transcription factors and a superior performance in identifying perturbed TFs compared to our previous.
See EnrichGT::infering_regulator_act() for help.
You should setReadable first
Like this: if your data is only contains gene symbols (like TP53, SIGLEC1, CD68…), please ignore this part.
enrichedObj |> setReadable(OrgDb = xxx,keyType = "ENTREZID") |> EnrichGT()
# orgDb can be org.Hs.eg.db, org.Mm.eg.db. Example
Generate EnrichGT_obj object
suppressMessages({
library(tidyverse)
library(gt)
library(clusterProfiler)
library(org.Hs.eg.db)
})
library(enrichplot) # for viewing enrichment result
data(geneList, package="DOSE")
gene <- names(geneList)[(geneList) > 1]
ego <- enrichGO(gene = gene,
universe = names(geneList),
OrgDb = org.Hs.eg.db,
ont = "BP",
pAdjustMethod = "BH",
pvalueCutoff = 0.5,
qvalueCutoff = 0.5,
readable = TRUE)
enrichGT_obj <- ego |> EnrichGT()
#> ℹ =====[SUGGESTION]=====
#> You are passing an object from GO Enrichment.
#> Please ensure that `obj |> clusterProfiler::simplify()` is executed, to pre-simplify result,
#> For better enriched result.
#> Loading required package: text2vec
#>
#> Attaching package: 'text2vec'
#> The following object is masked from 'package:BiocGenerics':
#>
#> normalize
Pathway Infering
pw_res<-enrichGT_obj |> infering_regulator_act(DB = "progeny", species = "human")
enrichplot::dotplot(pw_res)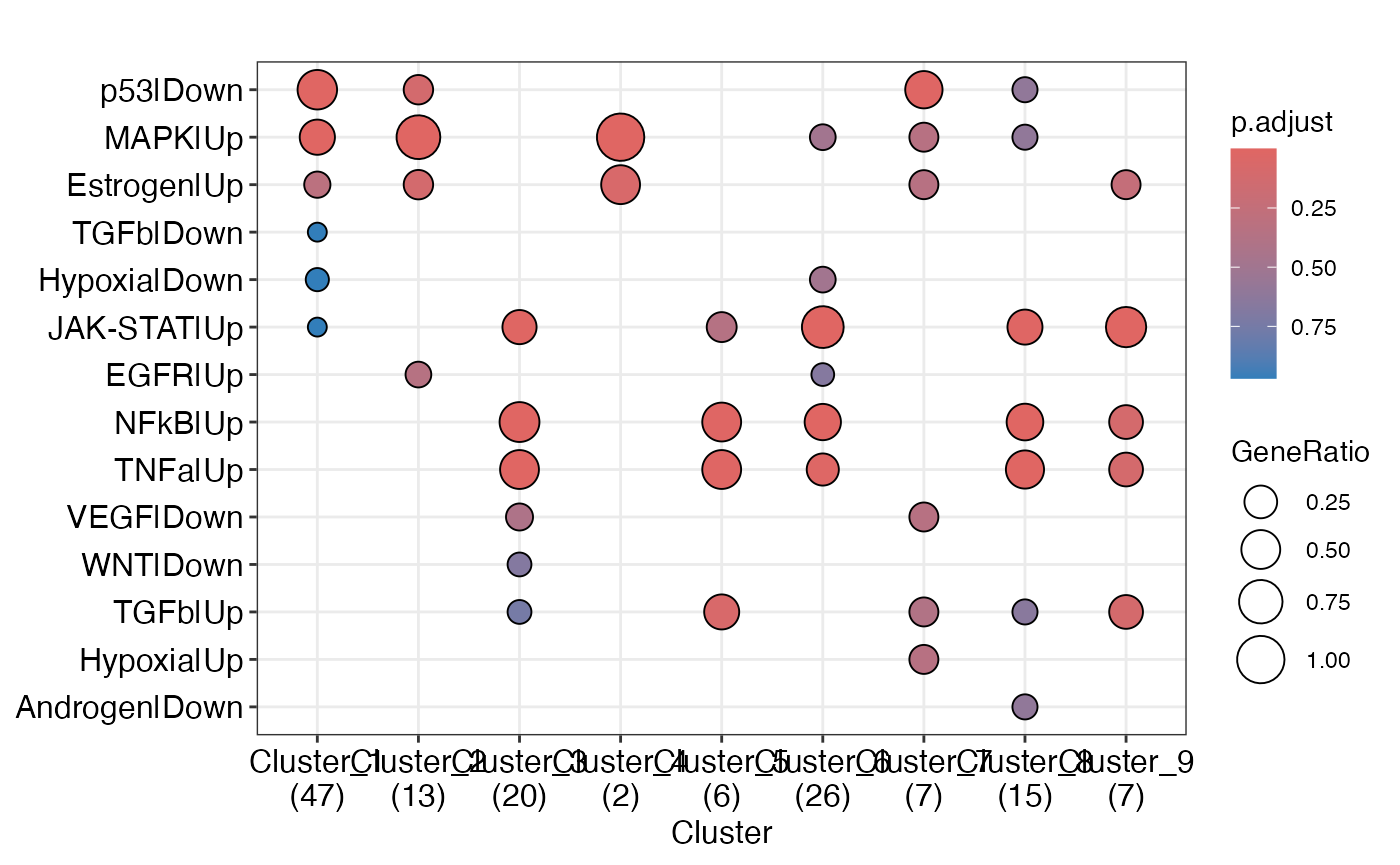
TF activity infering
tf_res<-enrichGT_obj |> infering_regulator_act(DB = "collectri", species = "human")
enrichplot::dotplot(tf_res)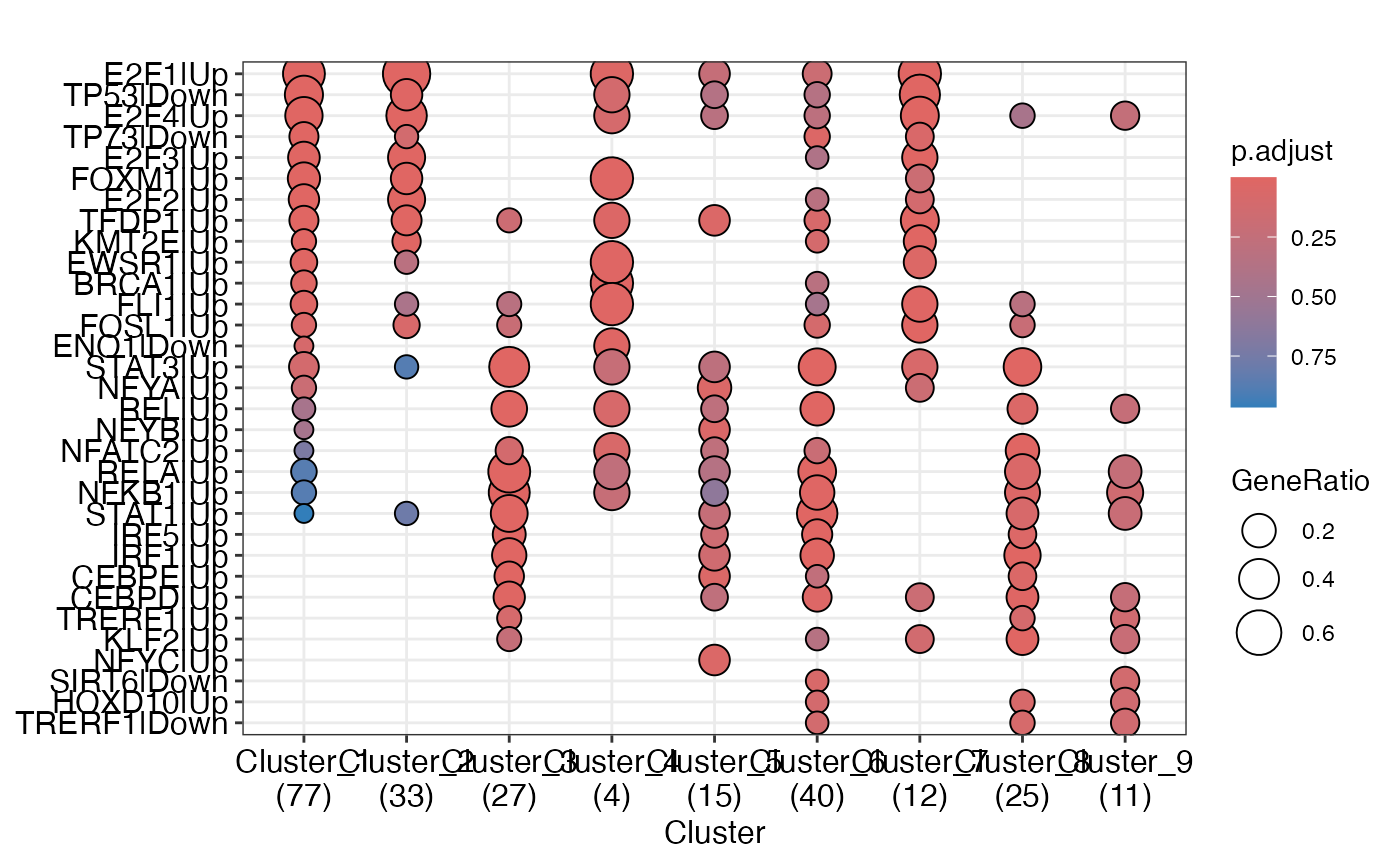
Session Info
sessioninfo::session_info()
#> ─ Session info ───────────────────────────────────────────────────────────────
#> setting value
#> version R version 4.3.3 (2024-02-29)
#> os macOS 15.0
#> system aarch64, darwin20
#> ui X11
#> language en
#> collate en_US.UTF-8
#> ctype en_US.UTF-8
#> tz Asia/Shanghai
#> date 2024-10-12
#> pandoc 3.1.11 @ /Applications/RStudio.app/Contents/Resources/app/quarto/bin/tools/aarch64/ (via rmarkdown)
#>
#> ─ Packages ───────────────────────────────────────────────────────────────────
#> package * version date (UTC) lib source
#> AnnotationDbi * 1.64.1 2023-11-02 [1] Bioconductor
#> ape 5.7-1 2023-03-13 [1] CRAN (R 4.3.0)
#> aplot 0.2.2 2023-10-06 [1] CRAN (R 4.3.1)
#> Biobase * 2.62.0 2023-10-26 [1] Bioconductor
#> BiocGenerics * 0.48.1 2023-11-02 [1] Bioconductor
#> BiocParallel 1.36.0 2023-10-26 [1] Bioconductor
#> Biostrings 2.70.2 2024-01-30 [1] Bioconductor 3.18 (R 4.3.2)
#> bit 4.0.5 2022-11-15 [1] CRAN (R 4.3.0)
#> bit64 4.0.5 2020-08-30 [1] CRAN (R 4.3.0)
#> bitops 1.0-7 2021-04-24 [1] CRAN (R 4.3.0)
#> blob 1.2.4 2023-03-17 [1] CRAN (R 4.3.0)
#> bslib 0.6.1 2023-11-28 [1] CRAN (R 4.3.1)
#> cachem 1.0.8 2023-05-01 [1] CRAN (R 4.3.0)
#> cli * 3.6.3 2024-06-21 [1] CRAN (R 4.3.3)
#> clusterProfiler * 4.10.1 2024-03-09 [1] Bioconductor 3.18 (R 4.3.3)
#> codetools 0.2-19 2023-02-01 [1] CRAN (R 4.3.3)
#> colorspace 2.1-0 2023-01-23 [1] CRAN (R 4.3.0)
#> cowplot 1.1.3 2024-01-22 [1] CRAN (R 4.3.1)
#> crayon 1.5.2 2022-09-29 [1] CRAN (R 4.3.0)
#> data.table 1.15.4 2024-03-30 [1] CRAN (R 4.3.1)
#> DBI 1.2.2 2024-02-16 [1] CRAN (R 4.3.1)
#> desc 1.4.3 2023-12-10 [1] CRAN (R 4.3.1)
#> digest 0.6.35 2024-03-11 [1] CRAN (R 4.3.1)
#> DOSE 3.28.2 2023-12-12 [1] Bioconductor 3.18 (R 4.3.2)
#> dplyr * 1.1.4 2023-11-17 [1] CRAN (R 4.3.1)
#> EnrichGT * 0.2.8.5 2024-10-12 [1] local
#> enrichplot * 1.22.0 2023-11-06 [1] Bioconductor
#> evaluate 0.23 2023-11-01 [1] CRAN (R 4.3.1)
#> fansi 1.0.6 2023-12-08 [1] CRAN (R 4.3.1)
#> farver 2.1.1 2022-07-06 [1] CRAN (R 4.3.0)
#> fastmap 1.1.1 2023-02-24 [1] CRAN (R 4.3.0)
#> fastmatch 1.1-4 2023-08-18 [1] CRAN (R 4.3.0)
#> fgsea 1.28.0 2023-10-26 [1] Bioconductor
#> float 0.3-2 2023-12-10 [1] CRAN (R 4.3.1)
#> forcats * 1.0.0 2023-01-29 [1] CRAN (R 4.3.0)
#> fs 1.6.3 2023-07-20 [1] CRAN (R 4.3.0)
#> generics 0.1.3 2022-07-05 [1] CRAN (R 4.3.0)
#> GenomeInfoDb 1.38.7 2024-03-09 [1] Bioconductor 3.18 (R 4.3.3)
#> GenomeInfoDbData 1.2.11 2024-03-18 [1] Bioconductor
#> ggforce 0.4.2 2024-02-19 [1] CRAN (R 4.3.1)
#> ggfun 0.1.4 2024-01-19 [1] CRAN (R 4.3.1)
#> ggplot2 * 3.5.0 2024-02-23 [1] CRAN (R 4.3.1)
#> ggplotify 0.1.2 2023-08-09 [1] CRAN (R 4.3.0)
#> ggraph 2.2.1 2024-03-07 [1] CRAN (R 4.3.1)
#> ggrepel 0.9.5 2024-01-10 [1] CRAN (R 4.3.1)
#> ggtree 3.10.1 2024-02-27 [1] Bioconductor 3.18 (R 4.3.2)
#> glue 1.7.0 2024-01-09 [1] CRAN (R 4.3.1)
#> GO.db 3.18.0 2024-03-18 [1] Bioconductor
#> GOSemSim 2.28.1 2024-01-20 [1] Bioconductor 3.18 (R 4.3.2)
#> graphlayouts 1.1.1 2024-03-09 [1] CRAN (R 4.3.1)
#> gridExtra 2.3 2017-09-09 [1] CRAN (R 4.3.0)
#> gridGraphics 0.5-1 2020-12-13 [1] CRAN (R 4.3.0)
#> gson 0.1.0 2023-03-07 [1] CRAN (R 4.3.0)
#> gt * 0.11.0 2024-07-09 [1] CRAN (R 4.3.3)
#> gtable 0.3.4 2023-08-21 [1] CRAN (R 4.3.0)
#> HDO.db 0.99.1 2024-03-18 [1] Bioconductor
#> highr 0.10 2022-12-22 [1] CRAN (R 4.3.0)
#> hms 1.1.3 2023-03-21 [1] CRAN (R 4.3.0)
#> htmltools 0.5.7 2023-11-03 [1] CRAN (R 4.3.1)
#> httr 1.4.7 2023-08-15 [1] CRAN (R 4.3.0)
#> igraph 2.0.3 2024-03-13 [1] CRAN (R 4.3.1)
#> IRanges * 2.36.0 2023-10-26 [1] Bioconductor
#> jquerylib 0.1.4 2021-04-26 [1] CRAN (R 4.3.0)
#> jsonlite 1.8.8 2023-12-04 [1] CRAN (R 4.3.1)
#> KEGGREST 1.42.0 2023-10-26 [1] Bioconductor
#> knitr 1.45 2023-10-30 [1] CRAN (R 4.3.1)
#> labeling 0.4.3 2023-08-29 [1] CRAN (R 4.3.0)
#> lattice 0.22-5 2023-10-24 [1] CRAN (R 4.3.3)
#> lazyeval 0.2.2 2019-03-15 [1] CRAN (R 4.3.0)
#> lgr 0.4.4 2022-09-05 [1] CRAN (R 4.3.0)
#> lifecycle 1.0.4 2023-11-07 [1] CRAN (R 4.3.1)
#> lubridate * 1.9.3 2023-09-27 [1] CRAN (R 4.3.1)
#> magrittr 2.0.3 2022-03-30 [1] CRAN (R 4.3.0)
#> MASS 7.3-60.0.1 2024-01-13 [1] CRAN (R 4.3.3)
#> Matrix 1.6-5 2024-01-11 [1] CRAN (R 4.3.3)
#> memoise 2.0.1 2021-11-26 [1] CRAN (R 4.3.0)
#> mlapi 0.1.1 2022-04-24 [1] CRAN (R 4.3.0)
#> munsell 0.5.0 2018-06-12 [1] CRAN (R 4.3.0)
#> nlme 3.1-164 2023-11-27 [1] CRAN (R 4.3.3)
#> org.Hs.eg.db * 3.18.0 2024-03-18 [1] Bioconductor
#> patchwork 1.2.0 2024-01-08 [1] CRAN (R 4.3.1)
#> pillar 1.9.0 2023-03-22 [1] CRAN (R 4.3.0)
#> pkgconfig 2.0.3 2019-09-22 [1] CRAN (R 4.3.0)
#> pkgdown 2.0.7 2022-12-14 [1] CRAN (R 4.3.0)
#> plyr 1.8.9 2023-10-02 [1] CRAN (R 4.3.1)
#> png 0.1-8 2022-11-29 [1] CRAN (R 4.3.0)
#> polyclip 1.10-6 2023-09-27 [1] CRAN (R 4.3.1)
#> proxy 0.4-27 2022-06-09 [1] CRAN (R 4.3.0)
#> purrr * 1.0.2 2023-08-10 [1] CRAN (R 4.3.0)
#> qvalue 2.34.0 2023-10-26 [1] Bioconductor
#> R6 2.5.1 2021-08-19 [1] CRAN (R 4.3.0)
#> ragg 1.3.0 2024-03-13 [1] CRAN (R 4.3.1)
#> RColorBrewer 1.1-3 2022-04-03 [1] CRAN (R 4.3.0)
#> Rcpp 1.0.12 2024-01-09 [1] CRAN (R 4.3.1)
#> RCurl 1.98-1.14 2024-01-09 [1] CRAN (R 4.3.1)
#> readr * 2.1.5 2024-01-10 [1] CRAN (R 4.3.1)
#> reshape2 1.4.4 2020-04-09 [1] CRAN (R 4.3.0)
#> RhpcBLASctl 0.23-42 2023-02-11 [1] CRAN (R 4.3.0)
#> rlang 1.1.3 2024-01-10 [1] CRAN (R 4.3.1)
#> rmarkdown 2.26 2024-03-05 [1] CRAN (R 4.3.1)
#> rsparse 0.5.2 2024-06-28 [1] CRAN (R 4.3.3)
#> RSQLite 2.3.5 2024-01-21 [1] CRAN (R 4.3.1)
#> rstudioapi 0.15.0 2023-07-07 [1] CRAN (R 4.3.0)
#> S4Vectors * 0.40.2 2023-11-25 [1] Bioconductor 3.18 (R 4.3.2)
#> sass 0.4.9 2024-03-15 [1] CRAN (R 4.3.1)
#> scales 1.3.0 2023-11-28 [1] CRAN (R 4.3.1)
#> scatterpie 0.2.1 2023-06-07 [1] CRAN (R 4.3.0)
#> sessioninfo 1.2.2 2021-12-06 [1] CRAN (R 4.3.0)
#> shadowtext 0.1.3 2024-01-19 [1] CRAN (R 4.3.1)
#> stringi 1.8.3 2023-12-11 [1] CRAN (R 4.3.1)
#> stringr * 1.5.1 2023-11-14 [1] CRAN (R 4.3.1)
#> systemfonts 1.1.0 2024-05-15 [1] CRAN (R 4.3.3)
#> text2vec * 0.6.4 2023-11-09 [1] CRAN (R 4.3.1)
#> textshaping 0.3.7 2023-10-09 [1] CRAN (R 4.3.1)
#> tibble * 3.2.1 2023-03-20 [1] CRAN (R 4.3.0)
#> tidygraph 1.3.1 2024-01-30 [1] CRAN (R 4.3.1)
#> tidyr * 1.3.1 2024-01-24 [1] CRAN (R 4.3.1)
#> tidyselect 1.2.1 2024-03-11 [1] CRAN (R 4.3.1)
#> tidytree 0.4.6 2023-12-12 [1] CRAN (R 4.3.1)
#> tidyverse * 2.0.0 2023-02-22 [1] CRAN (R 4.3.0)
#> timechange 0.3.0 2024-01-18 [1] CRAN (R 4.3.1)
#> treeio 1.26.0 2023-11-06 [1] Bioconductor
#> tweenr 2.0.3 2024-02-26 [1] CRAN (R 4.3.1)
#> tzdb 0.4.0 2023-05-12 [1] CRAN (R 4.3.0)
#> utf8 1.2.4 2023-10-22 [1] CRAN (R 4.3.1)
#> vctrs 0.6.5 2023-12-01 [1] CRAN (R 4.3.1)
#> viridis 0.6.5 2024-01-29 [1] CRAN (R 4.3.1)
#> viridisLite 0.4.2 2023-05-02 [1] CRAN (R 4.3.0)
#> withr 3.0.0 2024-01-16 [1] CRAN (R 4.3.1)
#> xfun 0.42 2024-02-08 [1] CRAN (R 4.3.1)
#> xml2 1.3.6 2023-12-04 [1] CRAN (R 4.3.1)
#> XVector 0.42.0 2023-10-26 [1] Bioconductor
#> yaml 2.3.8 2023-12-11 [1] CRAN (R 4.3.1)
#> yulab.utils 0.1.4 2024-01-28 [1] CRAN (R 4.3.1)
#> zlibbioc 1.48.0 2023-10-26 [1] Bioconductor
#>
#> [1] /Library/Frameworks/R.framework/Versions/4.3-arm64/Resources/library
#>
#> ──────────────────────────────────────────────────────────────────────────────